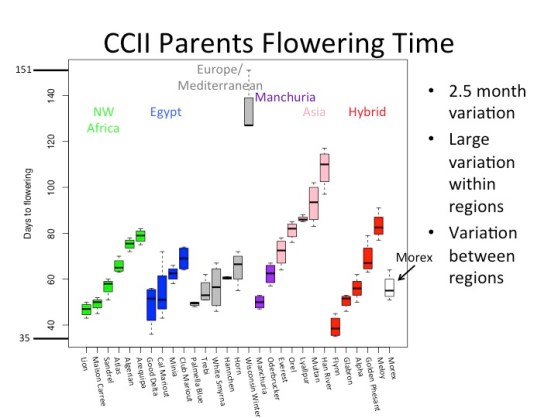
My previous research in the Koenig lab involves Evolutionary Genomics, focusing on the genomic changes associated with adaptation in barley (Hordeum vulgare). Specifically I used the Composite Cross II population, a near century long evolution experiment performed at UC Davis.
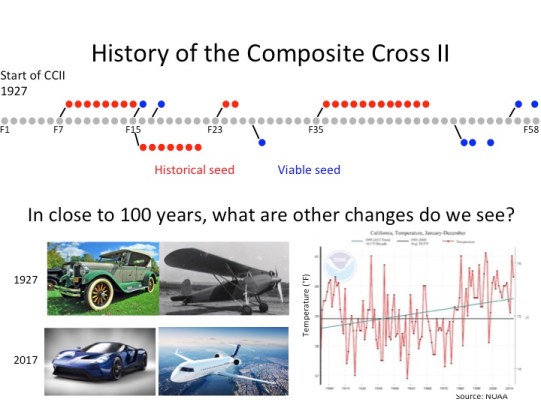
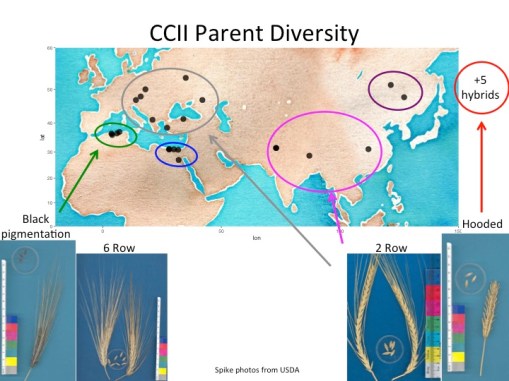
This population started out with reciprocal crosses of 28 landraces of barley which exhibited a wide range of the geographical and phenotypic diversity observed in barley. The progeny of the cross were grown in large generations without human selection for nearly 60 generations.
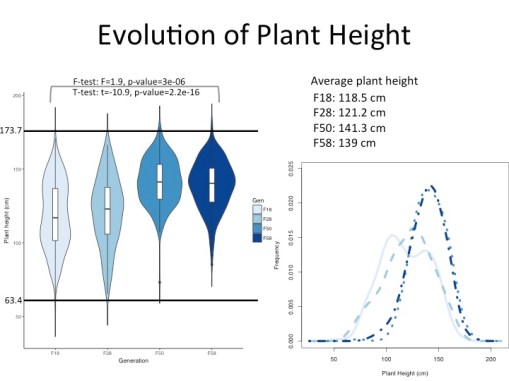
Using mostly common garden greenhouse conditions I am acquired phenotypes of the 28 parent landraces and progeny from four different generations to see if selection may be acting on traits of interest. Additionally, using a combination of RAD-Seq and Exome capture methods to generate SNPs for genotyping and future Genome Wide Association analyses to determine the genetic component of particular traits of interest. So far, we see that some traits appear to be impacted by purifying selection, while other traits look more like directional selection is acting on them.